Every living organism has its genome in the heart. If the cell is a computer, the genome is the software it executes. If we consider DNA as a cell-operated software, we can also use our own computers to analyze how it works with similar logic.DNA is not just an information store, it is a physical structure that can behave in a complex way. Genomes are incredibly complex machines with thousands of parts.
Although we know how some genes work today, it is not understood how many genes work together.
- Genetics sees DNA only as information, looks for patterns in knowledge, and investigates the relationships between genes and physical appearance
- Genomics sees the genome as a machine and tries to understand how its parts work together.
Genomic Data Science
Genomic Data Science is the application of methods such as statistics, machine learning, which are found in data science to genomic problems.
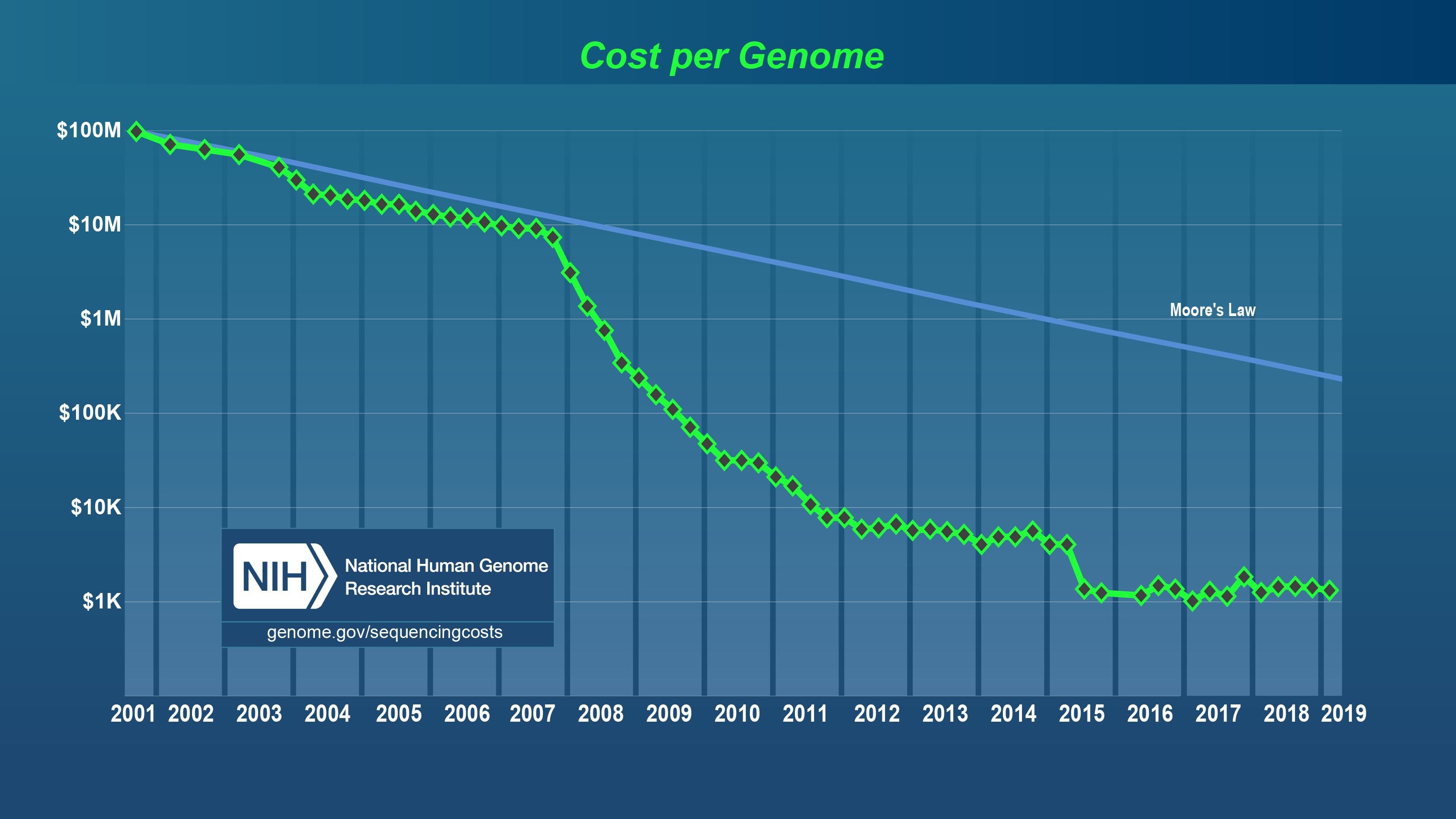
Thanks to next generation sequencing, we are able to sequence genomes faster, cheaper and successful. For example, the Human Genome Project was realized for $ 2.7 billion (in the past), and today you can sequence your genome for $ 1000.Therefore, a lot of genomic data can be found on the internet.
A living organism is more complex than any machine humans have ever produced. Solving this complexity is above the classical algorithms or the capacity of human understanding. Most genetic diseases and cancers occur through the interaction of multiple genes and contain genetic variation.
One of the most powerful weapons for analyzing data with incredibly complexity and data points is Deep Learning, classical methods assume a linear relationship between genes, which is not so.
With Genomic Data Science, personalized drug studies can be accelerated, or if you are bored, you can create a 3D version of their faces from genomes to detect people who have thrown cigarettes or chewing gum.
Life and DNA

Life from bacteria to whale works with similar principles. All living things are interconnected in the tree of life and most likely come from a single common ancestor.
DNA 4 is a long polymer consisting of repetition of the basic (A, T, G, C) base. Almost all the information on how to build the organism is stored here.
The code of this information itself and how it is processed (epigenetic) changes and evolves over time. DNA can carry information in an almost infinite combination.
Basic operation
Central Dogma
If DNA is software, proteins are the most important equipment. Proteins are small machines that do most of the work in the cell. When converting into proteins with the information in DNA, a molecule called mRNA is needed.
mRNA goes to ribosome and protein is synthesized by combining amino acids according to the information it contains.Ribosomes are organic 3D printers.
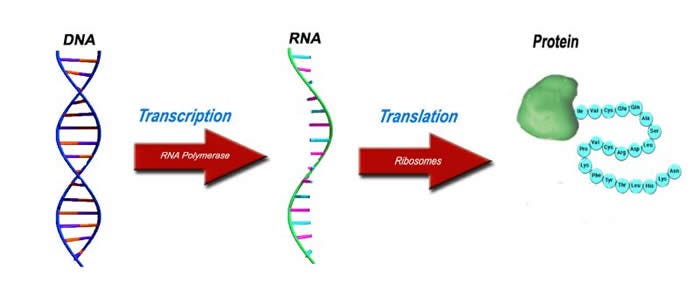
There is a one-way flow of information from DNA to protein and it works simply.
No matter how elegant this idea looks, it is incomplete according to our knowledge today.
The collapse of dogma and complexity
Now let’s examine how genomes really work.
-The DNA of eukaryotes is wrapped around proteins called histones to fit into the cell, the tightly packed portions cannot be read and the methylated portions become difficult to read. The mechanisms that regulate when DNA should be opened are not fully understood.
-There is no unidirectional flow of information from DNA to protein, and proteins can bind to DNA and act as regulators.
-mRNA knows which protein to synthesize but does not know when it should be synthesized.
-Transcription factors come into play here. Transcription factors bind to specific points in DNA and regulate the expression of the genes located nearby.
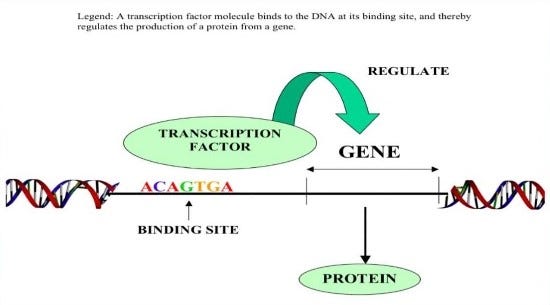
They owe differentiation of skin and nerve cells carrying the same genetic information to the regulation of gene expressions.

miRNA, siRNA, Riboswitches can participate in the regulation tasks.
Estimating the binding of the transcription factor (TF)
We found that a cell is very complex and difficult to work with classical methods, so we will practice deep learning techniques. We will use the HepG2 cell line and JunD TF as data. HepG2 is an immortal cell line derived from the liver tissue of a 15-year-old African-American child.

We choose the 22nd chromosome so that genomic data can be processed, this chromosome contains about 50 million base pairs, there are about 3 billion base pairs in man.The largest chromosome is the 1st and the smallest is 22nd.
If only 2 of the 3 billion base pairs (in a particular location) mutate, Cystic fibrosis or minor changes can lead to diseases such as Hemophilia, Color blindness, Muscular dystrophy.
Genomic data is stored in .FASTA or .FASTQ format. Data will have a similar view to book lines.
>chr22
NNNNNNNNNNNNNNNNNNNNNNNNNNN
TCCCAAATTGTGGAAGGAATGTACATTTGAC
Here we see the base sequences contained in a single chain of DNA, we can ask what the “N” symbol does in DNA, which means that it is not possible to decide which base is read in the sequencing process.
Next we have to convert the genomic information into a mathematical format that we can use. As we have seen, our data consists of A, T, G, C and N parts. 0,1,0] and N: [0.25,0.25,0.25,0.25] can be represented.
[[1., 0., 0., 0.],
[1., 0., 0., 0.],
[0., 1., 0., 0.], New chain
[0., 0., 1., 0.],
[0., 0., 0., 1.]]
Then we divide the chain into 101 length pieces. We know where the JunD protein can bind, so it becomes a problem of supervised learning.
For example, if JunD can be connected to the base arrays that we have broken down, we can add 1 as a label and 0 as a label.
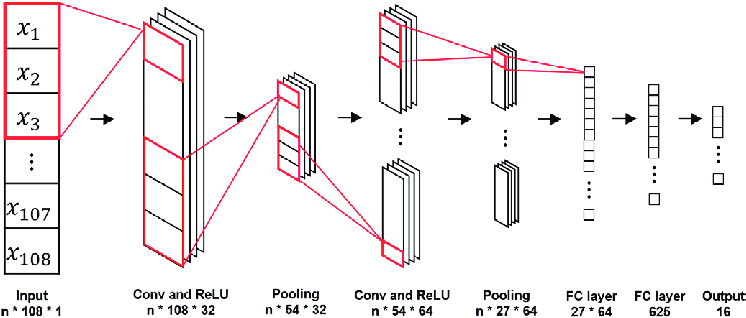
Since DNA is a string, it is one-dimensional. We can use different architectures. The most useful architecture for finding patterns is Evolutionary Neural Networks. They are generally used on images (2-dimensional information), and 1D CNNs can be used to process texts.
2D CNNs extract features from the image, learn edges, corners, color changes and more complex patterns.Similarly, we can learn filters that can extract attributes from text data. After our model has been trained, we can calculate how likely JunD protein will bind to DNA fragments of 101 base lengths.
Chromatin accessibility
There are many factors in the binding of JunD protein to DNA: accessibility, methylation, shape, the presence of other molecules in the environment.

Chromatin accessibility: Defines how accessible DNA is to molecules from outside. When DNA is tightly wrapped around histones, TFs or other molecules are inaccessible. Genes that are similarly silenced can also be packaged.
Accessibility of a region is not fixed, it changes over time. If we add chromatin accessibility information while training our model, we can make more successful predictions.
As a result: Machine learning can also improve the genomic field. Personalized drug and disease detection can be done by processing large genomic data. It also makes CRISPR gene editing technology, which is a different destructive technology, more effective.
— — Resources — –
https://www.oreilly.com/library/view/deep-learning-for/9781492039822/
Author

Ömer Özgür
DATA SCIENCE-AI TEAM